Probing Watson-Crick and Hoogsteen base pairing in duplex DNA using dynamic nuclear polarization solid-state NMR spectroscopy | PNAS
![Experiment 3 - 1, following data. (Must show working). [1 mark] Volume of culture:Calculate the - Studocu Experiment 3 - 1, following data. (Must show working). [1 mark] Volume of culture:Calculate the - Studocu](https://d20ohkaloyme4g.cloudfront.net/img/document_thumbnails/395e0496f7c3388f8526bae57bb22c15/thumb_1200_1698.png)
Experiment 3 - 1, following data. (Must show working). [1 mark] Volume of culture:Calculate the - Studocu

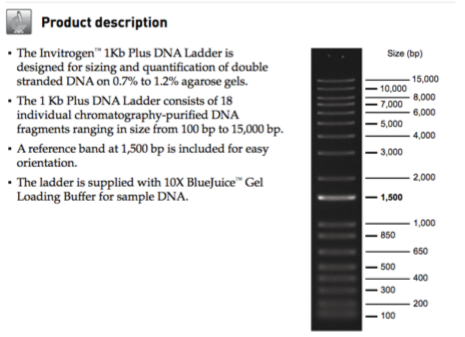
PCR products. Lane1, 1-kb molecular weight marker in base pairs; lane... | Download Scientific Diagram

Determining the mass of DNA fragments - BSCI 1510L Literature and Stats Guide - Research Guides at Vanderbilt University

anti–syn Unnatural Base Pair Enables Alphabet-Expanded DNA Self-Assembly | Journal of the American Chemical Society

SOLVED:The average molar mass of one base pair of nucleotides in DNA is approximately 600 g/mol. The spacing between successive base pairs is about 0.34 nm, and a complete turn in the
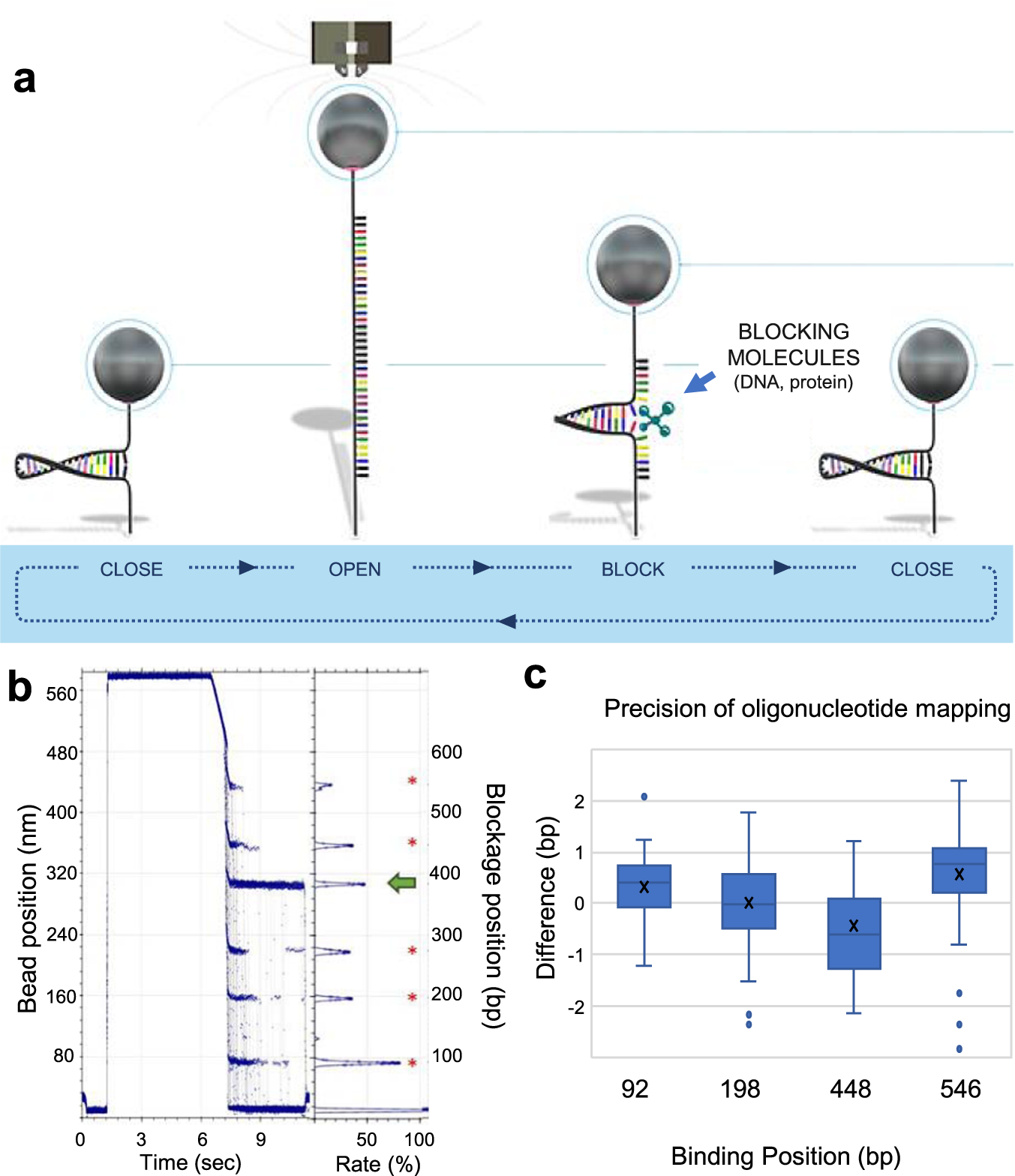
Detection of genetic variation and base modifications at base-pair resolution on both DNA and RNA | Communications Biology


.png?revision=1&size=bestfit&width=647&height=365)